
RODRÍGUEZ DE LA VEGA Ricardo C.
- Ecologie Systematique Evolution, Univ. Paris-Saclay - CNRS -AgroParisTech, Orsay, France
- Bioinformatics, Evolutionary genomics, Fungi, Proteomics
- recommender
Recommendation: 1
Reviews: 2
Recommendation: 1
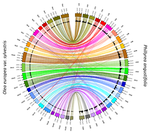
Genetic mapping of sex and self-incompatibility determinants in the androdioecious plant Phillyrea angustifolia
Identification of distinct YX-like loci for sex determination and self-incompatibility in an androdioecious shrub
Recommended by Tatiana Giraud and Ricardo C. Rodríguez de la Vega based on reviews by 2 anonymous reviewersA wide variety of systems have evolved to control mating compatibility in sexual organisms. Their genetic determinism and the factors controlling their evolution represent fascinating questions in evolutionary biology and genomics. The plant Phillyrea angustifolia (Oleaeceae family) represents an exciting model organism, as it displays two distinct and rare mating compatibility systems [1]: 1) males and hermaphrodites co-occur in populations of this shrub (a rare system called androdioecy), while the evolution and maintenance of purely hermaphroditic plants or mixtures of females and hermaphrodites (a system called gynodioecy) are easier to explain [2]; 2) a homomorphic diallelic self-incompatibility system acts in hermaphrodites, while such systems are usually multi-allelic, as rare alleles are advantageous, being compatible with all other alleles. Previous analyses of crosses brought some interesting answers to these puzzles, showing that males benefit from the ability to mate with all hermaphrodites regardless of their allele at the self-incompatibility system, and suggesting that both sex and self incompatibility are determined by XY-like genetic systems, i.e. with each a dominant allele; homozygotes for a single allele and heterozygotes therefore co-occur in natural populations at both sex and self-incompatibility loci [3].
Here, Carré et al. used genotyping-by-sequencing to build a genome linkage map of P. angustifolia [4]. The elegant and original use of a probabilistic model of segregating alleles (implemented in the SEX-DETector method) allowed to identify both the sex and self-incompatibility loci [4], while this tool was initially developed for detecting sex-linked genes in species with strictly separated sexes (dioecy) [5]. Carré et al. [4] confirmed that the sex and self-incompatibility loci are located in two distinct linkage groups and correspond to XY-like systems. A comparison with the genome of the closely related Olive tree indicated that their self-incompatibility systems were homologous. Such a XY-like system represents a rare genetic determination mechanism for self-incompatibility and has also been recently found to control mating types in oomycetes [6].
This study [4] paves the way for identifying the genes controlling the sex and self-incompatibility phenotypes and for understanding why and how self-incompatibility is only expressed in hermaphrodites and not in males. It will also be fascinating to study more finely the degree and extent of genomic differentiation at these two loci and to assess whether recombination suppression has extended stepwise away from the sex and self-incompatibility loci, as can be expected under some hypotheses, such as the sheltering of deleterious alleles near permanently heterozygous alleles [7]. Furthermore, the co-occurrence in P. angustifolia of sex and mating types can contribute to our understanding of the factor controlling their evolution [8].
References
[1] Saumitou-Laprade P, Vernet P, Vassiliadis C, Hoareau Y, Magny G de, Dommée B, Lepart J (2010) A Self-Incompatibility System Explains High Male Frequencies in an Androdioecious Plant. Science, 327, 1648–1650. https://doi.org/10.1126/science.1186687
[2] Pannell JR, Voillemot M (2015) Plant Mating Systems: Female Sterility in the Driver’s Seat. Current Biology, 25, R511–R514. https://doi.org/10.1016/j.cub.2015.04.044
[3] Billiard S, Husse L, Lepercq P, Godé C, Bourceaux A, Lepart J, Vernet P, Saumitou-Laprade P (2015) Selfish male-determining element favors the transition from hermaphroditism to androdioecy. Evolution, 69, 683–693. https://doi.org/10.1111/evo.12613
[4] Carre A, Gallina S, Santoni S, Vernet P, Gode C, Castric V, Saumitou-Laprade P (2021) Genetic mapping of sex and self-incompatibility determinants in the androdioecious plant Phillyrea angustifolia. bioRxiv, 2021.04.15.439943, ver. 7 peer-reviewed and recommended by Peer Community in Genomics. https://doi.org/10.1101/2021.04.15.439943
[5] Muyle A, Käfer J, Zemp N, Mousset S, Picard F, Marais GA (2016) SEX-DETector: A Probabilistic Approach to Study Sex Chromosomes in Non-Model Organisms. Genome Biology and Evolution, 8, 2530–2543. https://doi.org/10.1093/gbe/evw172
[6] Dussert Y, Legrand L, Mazet ID, Couture C, Piron M-C, Serre R-F, Bouchez O, Mestre P, Toffolatti SL, Giraud T, Delmotte F (2020) Identification of the First Oomycete Mating-type Locus Sequence in the Grapevine Downy Mildew Pathogen, Plasmopara viticola. Current Biology, 30, 3897-3907.e4. https://doi.org/10.1016/j.cub.2020.07.057
[7] Jay P, Tezenas E, Giraud T (2021) A deleterious mutation-sheltering theory for the evolution of sex chromosomes and supergenes. bioRxiv, 2021.05.17.444504. https://doi.org/10.1101/2021.05.17.444504
[8] Billiard S, López-Villavicencio M, Devier B, Hood ME, Fairhead C, Giraud T (2011) Having sex, yes, but with whom? Inferences from fungi on the evolution of anisogamy and mating types. Biological Reviews, 86, 421–442. https://doi.org/10.1111/j.1469-185X.2010.00153.x
Reviews: 2

Comparison of whole-genome assemblies of European river lamprey (Lampetra fluviatilis) and brook lamprey (Lampetra planeri)
Phased genomes suggest that L. fluviatilis and L. planeri are two ecotypes of the same species
Recommended by Samuel Abalde based on reviews by Ricardo C. Rodríguez de la Vega, Quentin Rougemont and 1 anonymous reviewerLampreys are the focus of intense research. Together with hagfishes, they form the Cyclostomata, the sister group of jawed vertebrates, and hence they are a key group for disentangling the early evolution of many vertebrate features (Shimel and Donoghue 2012; McCauley et al. 2015). Ecologically, lamprey species show a diverse array of life modes, including parasitic and non-feeding species, and inhabit freshwater and marine habitats or both (i.e. anadromous species; Docker and Potter 2019). One of these anadromous species, the sea lamprey (Petromyzon marinus), took advantage of man-made canals to invade the North American Great Lakes in the early 20th century, decimating many fish populations. Today, the control of these invasive populations is paramount for the survival of the region’s fishing industry (Ferreira-Martins et al. 2021). All these research avenues will benefit from the generation of new genomic data, an invaluable resource in evolutionary and conservation biology.
In this manuscript, Tørresen et al. (2025) present phased, chromosome-level assemblies from two lamprey species: the European river lamprey (Lampetra fluviatilis) and the brook lamprey (Lampetra planeri). These two genome assemblies are of high quality and will undoubtedly become a key resource in lamprey research. In particular, the authors showcase the potential of such genomes from two perspectives. First, comparing their assemblies to the already published genomes from P. marinus and another specimen of L. fluviatilis, they propose that lamprey genomes are highly conserved and display large syntenic blocks shared among species. Second, phylogenetic analyses and the annotation of SNPs suggest that L. fluviatilis and L. planeri should be considered two ecotypes of the same species complex, instead of two separate species. This might not be new for anyone knowledgeable in lamprey biology (Rougemont et al. 2017), but it is surprising given the distinct ecology of the two lampreys: L. fluviatilis is a parasitic, anadromous species, whereas L. planeri is a non-feeding, freshwater species.
In addition to the biological significance of this manuscript, I would like to acknowledge the robustness of the analytical approaches. These genomes were assembled and annotated following two pipelines recently developed at EBP-Nor, the Norwegian initiative of the Earth BioGenome Project (EBP). These pipelines are designed to be an easy-to-use, end-to-end solution for genomic analyses and are likely to become a standard for the EBP and European Reference Genome Atlas initiatives. There can be no better evidence of their effectiveness than these two phased, chromosome-level, highly complete genome assemblies.
References
Docker MF, Potter IC (2019) Life history evolution in lampreys: Alternative migratory and feeding types. In: Docker M (ed) Lampreys: Biology, Conservation and Control. Fish & Fisheries Series, vol 38. Springer, Dordrecht. https://doi.org/10.1007/978-94-024-1684-8_4
Ferreira-Martins D, Champer J, McCauley DW, Zhang Z, Docker MF (2021) Genetic control of invasive sea lamprey in the Great Lakes. Journal of Great Lakes Research, 47, S764-S775. https://doi.org/10.1016/j.jglr.2021.10.018
McCauley DW, Docker MF, Whyard S, Li W (2015) Lampreys as diverse model organisms in the genomics era. BioScience, 65(11), 1046-1056. https://doi.org/10.1093/biosci/biv139
Rougemont Q, Gagnaire PA, Perrier C, Genthon C, Besnard AL, Launey S, Evanno G (2017) Inferring the demographic history underlying parallel genomic divergence among pairs of parasitic and nonparasitic lamprey ecotypes. Molecular Ecology, 26(1), 142-162. https://doi.org/10.1111/mec.13664
Shimeld SM, Donoghue PC (2012) Evolutionary crossroads in developmental biology: cyclostomes (lamprey and hagfish). Development, 139(12), 2091-2099. https://doi.org/10.1242/dev.074716
Tørresen OK, Garmann-Aarhus B, Hoff SNK, Jentoft S, Svensson M, Schartum E, Tooming-Klunderud A, Skage M, Krabberød A, Vøllestad LA, Jakobsen KS (2025) Comparison of whole-genome assemblies of European river lamprey (Lampetra fluviatilis) and brook lamprey (Lampetra planeri). bioRxiv, ver. 5 peer-reviewed and recommended by PCI Genomics https://doi.org/10.1101/2024.12.06.627158
Nucleosome patterns in four plant pathogenic fungi with contrasted genome structures
Genome-wide chromatin and expression datasets of various pathogenic ascomycetes
Recommended by Sébastien Bloyer and Romain Koszul based on reviews by Ricardo C. Rodríguez de la Vega and 1 anonymous reviewerPlant pathogenic fungi represent serious economic threats. These organisms are rapidly adaptable, with plastic genomes containing many variable regions and evolving rapidly. It is, therefore, useful to characterize their genetic regulation in order to improve their control. One of the steps to do this is to obtain omics data that link their DNA structure and gene expression.
In this paper, Clairet et al. (2022) studied the nucleosome positioning and gene expression of four plant pathogenic ascomycete species (Leptosphaeria maculans, Leptosphaeria maculans 'lepidii', Fusarium graminearum, Botrytis cinerea). The genomes of these species contain different compositions of transposable elements (from 4 to 30%), and present an equally variable compartmentalization. The authors established MNAse-seq and RNA-seq maps of these genomes in axenic cultures. Thanks to an ad-hoc tool allowing the visualization of MNA-seq data in combination with other "omics" data, they were able to compare the maps of the different species between them and to study different types of correlation. This tool, called MSTS for "MNase-Seq Tool Suite", allows for example to perform limited analyses on certain genetic subsets in an ergonomic way.
In the fungi studied, nucleosomes are positioned every 161 to 172 bp, with intra-genome variations such as AT-rich regions but, surprisingly, particularly dense nucleosomes in the Lmb genome. The authors discuss the differences between these organisms with respect to this nucleosome density, the expression profile, and the structure and transposon composition of the different genomes. These data and insights thus represent interesting resources for researchers interested in the evolution of ascomycete genomes and their adaptation. For this, and for the development of the MSTS tool, we recommend this preprint.
References
Clairet C, Lapalu N, Simon A, Soyer JL, Viaud M, Zehraoui E, Dalmais B, Fudal I, Ponts N (2022) Nucleosome patterns in four plant pathogenic fungi with contrasted genome structures. bioRxiv, 2021.04.16.439968, ver. 4 peer-reviewed and recommended by Peer Community in Genomics. https://doi.org/10.1101/2021.04.16.439968