
Genome-wide chromatin and expression datasets of various pathogenic ascomycetes
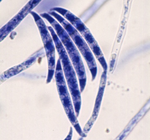
Nucleosome patterns in four plant pathogenic fungi with contrasted genome structures
Abstract
Recommendation: posted 04 July 2022, validated 13 July 2022
Bloyer, S. and Koszul, R. (2022) Genome-wide chromatin and expression datasets of various pathogenic ascomycetes. Peer Community in Genomics, 100014. https://doi.org/10.24072/pci.genomics.100014
Recommendation
Plant pathogenic fungi represent serious economic threats. These organisms are rapidly adaptable, with plastic genomes containing many variable regions and evolving rapidly. It is, therefore, useful to characterize their genetic regulation in order to improve their control. One of the steps to do this is to obtain omics data that link their DNA structure and gene expression.
In this paper, Clairet et al. (2022) studied the nucleosome positioning and gene expression of four plant pathogenic ascomycete species (Leptosphaeria maculans, Leptosphaeria maculans 'lepidii', Fusarium graminearum, Botrytis cinerea). The genomes of these species contain different compositions of transposable elements (from 4 to 30%), and present an equally variable compartmentalization. The authors established MNAse-seq and RNA-seq maps of these genomes in axenic cultures. Thanks to an ad-hoc tool allowing the visualization of MNA-seq data in combination with other "omics" data, they were able to compare the maps of the different species between them and to study different types of correlation. This tool, called MSTS for "MNase-Seq Tool Suite", allows for example to perform limited analyses on certain genetic subsets in an ergonomic way.
In the fungi studied, nucleosomes are positioned every 161 to 172 bp, with intra-genome variations such as AT-rich regions but, surprisingly, particularly dense nucleosomes in the Lmb genome. The authors discuss the differences between these organisms with respect to this nucleosome density, the expression profile, and the structure and transposon composition of the different genomes. These data and insights thus represent interesting resources for researchers interested in the evolution of ascomycete genomes and their adaptation. For this, and for the development of the MSTS tool, we recommend this preprint.
References
Clairet C, Lapalu N, Simon A, Soyer JL, Viaud M, Zehraoui E, Dalmais B, Fudal I, Ponts N (2022) Nucleosome patterns in four plant pathogenic fungi with contrasted genome structures. bioRxiv, 2021.04.16.439968, ver. 4 peer-reviewed and recommended by Peer Community in Genomics. https://doi.org/10.1101/2021.04.16.439968
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
This work was financially supported by the Plant Health and Environment division of the French National Institute for Agricultural Research (TACTIC Project AAP 2014). J. L. Soyer was funded by a “Contrat Jeune Scientifique” grant from INRAE. The BIOGER Unit benefits from the support of Saclay Plant Sciences-SPS (ANR-17-EUR-0007).
Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.1101/2021.04.16.439968
Version of the preprint: 1
Author's Reply, 18 Feb 2022
Decision by Sébastien Bloyer, posted 11 Jun 2021
Based on the comments of the two reviewers, I recommend that the authors correct and respond to the main suggestions highlighted in the reviews.
All the best,
Reviewed by anonymous reviewer 1, 04 Jun 2021
Reviewed by Ricardo C. Rodríguez de la Vega, 11 Jun 2021
Comments on Clairet et al., "Nucleosome patterns in four plant pathogenic..."
in this preprint authors present the analysis of nucleosome occupancy on four plant pathogenic fungi and discuss several factors that might underlie the observed patterns. Authors developed a freely available bioinformatics workflow to analyse and plot nucleosome phasing data that might be of interest to other researchers. The MNase-seq datasets could be reused for larger comparative nucleosome occupancy studies. I think, however, that the preprint is spare on technical details and overinterpret the results in some parts. Below I detail my general comments, point to specific aspects that I would like to see addressed and finish with some minor corrections/clarifications. I hope authors find the comments useful and are willing to address the points raised.
General
1. The bioinformatics workflow is hardly described at all, thus leaving out some important details. These include both plotting parameters used in the different parts of MSTS and statistical choices. For instance,
- a) in the methods I couldn't find how nucleosome signal intensity was "normalized", but in figures' legends it is stated that these are z-score normalized, is this the only option available in MSTS? Why are there no positive normalized nucleosome signal values on figure 3A?
- b) as far as I can tell MSTS reports the standard deviation of inter-nucleosome distances. In the preprint standard errors are reported, for me it is not clear the reason behind, what was the sample size used to transform SD to SE? Authors interpret the minute differences in nucleosome packing between SNP rich and SNP poor region as evidence of "increased frequencies of transient nucleosome positioning events in F. graminearum fast evolving polymorphic islands'' (page 7, second paragraph), but no statistical test was applied and if the value oscillates 1 bp around the mean, the difference becomes irrelevant.
- c) in my reading "conserved genes" would have almost no variance on nucleosome positioning at the first ATG codon. I actually wonder if the variance in +1 nucleosome and preceding NDR could be due to misprediction of starting codons, would expect the most curated gene models would show a lesser effect as a larger number of annotated ATG would indeed be the starting codon) It seems prediction holds ( see figures 6 and 7)
- d) it is not clear to me why the nucleosome signal intensity was not normalized in figure 8, neither why there's no estimation of the variance for the nucleosome depleted region (NDR) before the transcription start site/ATG codon. I couldn't find the sample sizes for the different expression categories, but as the average approaches the true mean for larger sample sizes, I would expect the different expression categories would have different accuracies regarding the actual position of the NDR. If low expressions are a minority, this could explain the erratic period seen across expression categories in figure 8 and the neat periods in figure 7. As of now, I can't phantom how the neat period after the ATG in figure 7 could be generated by the sum of waves in figure 8.
- e) inter nucleosome distances are more homogeneous with respect to the transcription start site (TSS) than with respect to the predicted translation start site, could the authors offer an explanation? could it be that the first methionine codon is mispredicted as the translation start site? For instance, comparing figures 8C with 8E and 8D with 8F it is clear that nucleosome phasing in Botrytis and Fusarium is much more homogeneous before and after the TSS than with respect to the predicted start of the sequence, how come? Either, there would be two NDR in these species, one at TSS and one at the start codon, or it is the same NDR and the nucleosome at +1 is either less well fixed or ATG is mispredicted (in this case the distance between the TSS and the ATG would oscillate in a narrow range). This is also relevant for the discussion on whether or not the position of NDR is less strict with respect to the TSS.
2. Some figures appear as a patchwork, with mixed framed and not-framed plots (e.g. figures 3 and 4), clipped ranges (e.g. figure 3C), inconsistent legend and inset position. I also think the choice of presenting dinucleotide frequencies in different scales is misleading (e.g. representation belies that values for dinucleotides involving C and G are lower in three species, in Botrytis values ranged from 0.176 to 0.181 for dinucleotides involving C and G but from 0.313 to 0.321 for dinucleotides involving A and T). Why the Rorschach-like plots (figure a5) are not strictly symmetrical? In general smoothed lines in phaseogram plots should be made thicker and the chromosome boundaries in figures 2 and 3 represented as gaps rather than vertical lines.
Specific points
- Please make the life of reviewers easier, mark the line numbers on the manuscript.
- Could authors comment on whether they expect nucleosome profiles would change depending on the physiological and metabolic state of the sequenced fungi?
- I think authors overplay the "genome compartalization" card, I reckon there are as many phytopathogenic fungi with largely homogeneous genomes (in terms of mobile elements for instance) as there are with compartmentalized genomes (e.g. Torrres et al., 2020 doi:10.1016/j.fbr.2020.07.001)
- I don't understand what authors meant by saying that "nucleosome positioning and occupancies are subjected to evolution" (p1, abstract) and "NDRs positions and intensities are subjected to evolution" (p10, third paragraph). Everything in biology is of course subjected to evolution, but what the authors showed here is that nucleosome phasing varies depending on genome context and species. Whether these genome contexts and species specific patterns are generalizable is not addressed in this preprint.
- State the distance between the known TSS and the predicted translation start site for Botrytis and Fusarium.
- What's the average gene size in the genomes? For Fusarium a drop on nucleosome occupancy over the gene body is clear on figure 7C.
Minor points and proofreading (suggested changes in upper case)
p1, abstract: spell out MSTS, e.g. "we developed the tool MSTS (FOR MNASE-SEQ TOOL SUITE)" or "we DEVELOPED MNASE-SEQ TOOL SUITE (MSTS)"
p1, introduction: comparing fungi (an entire Kingdom) with Insects (a class within the kingdom Animalia) is unfair. I think equating number of named species with biodiversity is misleading for a number of factors (size of the community studying them, research effort, inconsistent species delimitations practices across disciplines, etc)
p2, first paragraph: "important damages in agriculture, human health, and THE environment" ... "and facilitate infection. EFFECTORS CAN BE small proteins"
p2, first paragraph: Add references to the statement "Upon plant infection, fungi undergo a tightly controlled transcriptional reprogramming..."
p2, first paragraph: how can we speak of "plastic regions" for genomes with "overall large proportion of TE evenly distributed throughout the genome"?
p2, first paragraph: please cite the "several recent studies (pointing) out the potential role of chromatin remodelling" instead of the 6-10 years old reviews
p2, second paragraph: "hemiascomycetous yeasts. NO comparative genome-wide analyses"
p2, third paragraph: "four different plant pathogenic fungi ASCOMYCETES showing"
p3, second paragraph: "micrococcal nuclease digestion of mono-nucleosomes COUPLED WITH HIGH-THROUGHPUT SEQUENCING (MAINE-seq or MNase-seq)"
p3, second paragraph: not sure to which "contrasted media" authors refer to. Only one media per species was used.
p5, second paragraph and elsewhere: homogenize the abbreviation of use of micrococcal nuclease digestion of mono-nucleosomes coupled with high-throughput sequencing
p5, third paragraph: can you comment on whether the estimated linker lengths (14 to 25 bp) are within what's known for other fungi?
p6, second paragraph: "plants and other higher eukaryotes'', really? "higher" to what?
p7, second paragraph: "regions equally packed with nucleosomes are interspaced with AREAS with lower density"
p8, second paragraph: "previously described ~10 bp-periodicities", previously described where?
p10, third paragraph: "when the analysis is restricted to CONSERVED fungal genes". Mind what "culture conditions" are not a factor here as they are linked to the species.
p21, figure 4 legend: add the reference of Laurent et al. (2017) to explain how "TE and AT-rich regions" were defined










