
The number of Y chromosomes is positively associated with transposable element expression in humans, in line with the toxic Y hypothesis
 based on reviews by 3 anonymous reviewers
based on reviews by 3 anonymous reviewers
Transposable element expression with variation in sex chromosome number supports a toxic Y effect on human longevity
Abstract
Recommendation: posted 16 June 2024, validated 26 June 2024
Fiston-Lavier, A.-S. (2024) The number of Y chromosomes is positively associated with transposable element expression in humans, in line with the toxic Y hypothesis. Peer Community in Genomics, 100293. https://doi.org/10.24072/pci.genomics.100293
Recommendation
The study of human longevity has long been a source of fascination for scientists, particularly in relation to the genetic factors that contribute to differences in lifespan between the sexes. One particularly intriguing area of research concerns the Y chromosome and its impact on male longevity. The Y chromosome expresses genes that are essential for male development and reproduction. However, it may also influence various physiological processes and health outcomes. It is therefore of great importance to investigate the impact of the Y chromosome on longevity. This may assist in elucidating the biological mechanisms underlying sex-specific differences in aging and disease susceptibility. As longevity research progresses, the Y chromosome's role presents a promising avenue for elucidating the complex interplay between genetics and aging.
Transposable elements (TEs), often referred to as "jumping genes", are DNA sequences that can move within the genome, potentially causing mutations and genomic instability. In young, healthy cells, various mechanisms, including DNA methylation and histone modifications, suppress TE activity to maintain genomic integrity. However, as individuals age, these regulatory mechanisms may deteriorate, leading to increased TE activity. This dysregulation could contribute to age-related genomic instability, cellular dysfunction, and the onset of diseases such as cancer. Understanding how TE repression changes with age is crucial for uncovering the molecular underpinnings of aging (De Cecco et al. 2013; Van Meter et al. 2014).
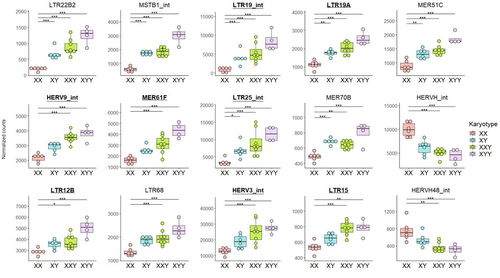
The lower recombination rates observed on Y chromosomes result in the accumulation of TE insertions, which in turn leads to an enrichment of TEs and potentially higher TE activity. To ascertain whether the number of Y chromosomes is associated with TE activity in humans, Teoli et al. (2024) studied the TE expression level, as a proxy of the TE activity, in several karyotype compositions (i.e. with differing numbers of Y chromosomes). They used transcriptomic data from blood samples collected in 24 individuals (six females 46,XX, six males 46,XY, eight males 47,XXY and four males 47,XYY). Even though they did not observe a significant correlation between the number of Y chromosomes and TE expression, their results suggest an impact of the presence of the Y chromosome on the overall TE expression. The presence of Y chromosomes also affected the type (family) of TE present/expressed. To ensure that the TE expression level was not biased by the expression of a gene in proximity due to intron retention or pervasive intragenic transcription, the authors also tested whether the TE expression variation observed between the different karyotypes could be explained by gene (i.e. here non-TE gene) expression.
As TE repression mechanisms are known to decrease over time, the authors also tested whether TE repression is weaker in older individuals, which would support a compelling link between genomic stability and aging. They investigated the TE expression differently between males and females, hypothesizing that old males should exhibit a stronger TE activity than old females. Using selected 45 males (47,XY) and 35 females (46,XX) blood samples of various ages (from 20 to 70) from the Genotype-Tissue Expression (GTEx) project, the authors studied the effect of age on TE expression using 10-year range to group the study subjects. Based on these data, they fail to find an overall increase of TE expression in old males compared to old females.
Notwithstanding the small number of samples, the study is well-designed and innovative, and its findings are highly promising. It marks an initial step towards understanding the impact of Y-chromosome ‘toxicity’ on human longevity. Despite the relatively small sample size, which is a consequence of the difficulty of obtaining samples from individuals with sex chromosome aneuploidies, the results are highly intriguing and will be of interest to a broad range of biologists.
References
De Cecco M, Criscione SW, Peckham EJ, Hillenmeyer S, Hamm EA, Manivannan J, Peterson AL, Kreiling JA, Neretti N, Sedivy JM (2013) Genomes of replicatively senescent cells undergo global epigenetic changes leading to gene silencing and activation of transposable elements. Aging Cell, 12, 247–256. https://doi.org/10.1111/acel.12047
Teoli J, Merenciano M, Fablet M, Necsulea A, Siqueira-de-Oliveira D, Brandulas-Cammarata A, Labalme A, Lejeune H, Lemaitre J-F, Gueyffier F, Sanlaville D, Bardel C, Vieira C, Marais GAB, Plotton I (2024) Transposable element expression with variation in sex chromosome number supports a toxic Y effect on human longevity. bioRxiv, ver. 5 peer-reviewed and recommended by Peer Community in Genomics. https://doi.org/10.1101/2023.08.03.550779
Van Meter M, Kashyap M, Rezazadeh S, Geneva AJ, Morello TD, Seluanov A, Gorbunova V (2014) SIRT6 represses LINE1 retrotransposons by ribosylating KAP1 but this repression fails with stress and age. Nature Communications, 5, 5011. https://doi.org/10.1038/ncomms6011
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
ANR-20-CE02-0015
Reviewed by anonymous reviewer 2, 30 May 2024
I have read the new version of the manuscript and the authors have addressed most of my comments.
The paragraph L439-461 is important as it provides more nuance on the way TE expression may be associated with ageing. I would recommend that this appears more clearly in the Introduction which is still mostly revolving around the idea of somatic mutation.
Section Data, scripts, code, and supplementary information availability
I note that the scripts will be "made available upon request". However there is a Github https://github.com/teolijo/longevitY/tree/main which could be edited to add a few pieces of information. I would recommend that the ling appears in the manuscript to help readers who may be interested in reproducing or testing the pipeline.
https://doi.org/10.24072/pci.genomics.100293.rev21
Reviewed by anonymous reviewer 3, 02 May 2024
Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.1101/2023.08.03.550779
Version of the preprint: 1
Author's Reply, 30 Mar 2024
We posted the final formatted version on the preprint server (no track change mode).
Please find the author's reply to reviewers attached as a PDF file.
Best regards.
Decision by Anna-Sophie Fiston-Lavier , posted 16 Nov 2023, validated 22 Nov 2023
, posted 16 Nov 2023, validated 22 Nov 2023
In this article, Teoli et al analyzed the variation in the expression level of transposable element (TE) insertions in RNAseq datasets from individuals with various numbers of sex chromosomes in order to test the toxic effect of Y on human lifespan, a fascinating but still controversial subject.
Specifically, they tested two predictions. These predictions suggest that the genomes of an older man should harbor more TEs compared to those of an older woman, due to the toxic effect of the Y chromosome and less efficient epigenetic regulation with age. In addition, they also tested whether TE expression varies with the number of Y chromosome using RNA resequencing data from 25 blood samples from individuals with different karyotypic compositions (46,XX, 46,XY, 47,XXY and 47,XYY).
However, the two reviewers noted several points which it seems important to address. One important point concerns the methodology of part 1, in which you present the analysis of a selected subset of GTEx data. Although reviewer 1 appreciates your honesty about your hindsight on the results of this part, the justification of the choice of data and filters used is missing. Reviewer 2 also suggested to look at the TEs that are expected to be more likely involved in somatic mutations.
As I believe this is a relevant topic that will be of interest to the community, I encourage the authors to respond carefully to the various points highlighted by the reviewers, and to resubmit their article.
Reviewed by anonymous reviewer 1, 02 Oct 2023
In this manuscript, Teoli and colleagues analyse expression data from human individuals to test the hypothesis of the Toxic Y, which could be involved in explaining the shorter lifespan of males compared to females in our species and many others. They used two independent datasets: 1. the blood RNA-seq data from the Genotype-Tissue Expression (GTEx) project and 2. a blood RNA-seq dataset from several individuals with different karyotypes generated by the authors. Analyses of dataset 1 provided some support for the prediction that TEs reactivate in old individuals, particularly in males. However, the structure of the data prevented the author from drawing firm conclusions and the results are quite noisy. The analyses of the second dataset provide better support for the second prediction, which suggests that TE expression is associated with the number of Y chromosomes.
Overall, I think the paper is concise, clear and well written. The analyses seem to have been carried out carefully and I really appreciate that the methods used are described in detail. I also appreciate that the authors do not overstate their findings and are honest about the limitations of their study, particularly the small sample size. The results are very interesting and should be of interest to a wide range of biologists.
My only major concern is with the analyses of dataset 1. In short, the authors greatly reduce the size of the genotype-tissue expression dataset by applying very strict filtering. This strong filtering is not properly justified and results in a very small dataset to analyse. As RNA-seq data are generally very noisy, this small dataset prevents the authors from drawing conclusions. I'm not sure that this strong filtering is necessary, and it should be better justified. For example, in "To reduce data heterogeneity and get closer to the individuals we sampled in parallel to constitute our dataset", I do not understand why you need a dataset "close" to the individuals you sampled: you do not analyse the two datasets together, but separately. Reducing the heterogeneity of the data only makes sense if that heterogeneity is biasing your results, which is not necessarily the case. Removing half of the dataset by eliminating non-white or Latino/Hispanic individuals seems to me to be too strong a filter and is not justified. It greatly reduces your statistical power. The same rationale applies to the other filters, such as excluding people with viruses, cancer or dementia. I understand how these disorders might affect gene or TE expression, but I'm not sure how this should affect the putative toxic Y effect. I think you should try to re-analyse this dataset with very minimal or no filtering, possibly including these inter-sample differences (ethnicity, disorders, etc.) as covariates in your statistical model when comparing TE expression between sexes and age groups. Alternatively, you should properly justify why these filters are necessary (e.g. strong interactions between ethnicity and age class in TE expression).
Other comments:
Abstract:
“Lifespan differences between sexes is a puzzling question”. For me, a difference is not a question.
“a toxic genomic impact in this trait” What is the trait ? Male lifespan ?
L59: Should be “Z-linked”, no ?
L65-68: The use of “is caused by” and ‘is still controversial” seems a bit contradictory to me.
L69-80: I think you should quickly define 46,XX, 46,XY, 47XXY, 47,XYY. It may not be clear to all readers what this means.
L104: “female-biased protein-coding genes”. A bit hard to read and understand at the beginning. Maybe it could be reformulated as, e.g. “genes overexpressed in females”?
L104: “most” → How many ?
L106: “It is indeed well-known that the extra-X chromosomes in 47,XXY or other karyotypes (e.g triple X) are inactivated” Not clear. The previous sentences refer to XX and XY samples, not XXY. The connection between this sentence and the previous ones should be explained. Does the sentence “We also found that most (13/20, 65%) male-biased protein-coding genes are Y-linked genes, and that most female-biased protein-coding genes are known X chromosome inactivation escapees” discuss only results comparing XX and XY or also other genotypes ?
Fig 1 legend: I think there is an error. I think it should be : “ […] and less than 0.01 (**) for 47,XYY samples compared to ##46,XX samples## (P = 0.0095) and near 0.05 for 47,XYY samples compared to 46,XY samples (P = 0.067) using Wilcoxon test”
Fig S9: What do represent the blue and grey arrows ?
Fig S11: What is the x axis ?
L120-175: I think it is important to write and discuss that an important observation is that XXY and XYY individuals have more TE expression than XX and XY individuals, which suggests that the number of sex chromosomes is a major determinant of TE expression (more than the number of Y chromosomes). The difference in TE expression between XXY and XYY is small and not significant. So the claim that you show an over-expression of TE is related to the number of Y chromosomes in the karyotype seems a bit too strong to me.
L155-156: ”Strong change” & “Clear trend”. Considering that these changes and trends are not statistically significant, I think you should tone down this sentence a bit.
https://doi.org/10.24072/pci.genomics.100293.rev11










