
A flexible and reproducible pipeline for long-read assembly and evaluation
CulebrONT: a streamlined long reads multi-assembler pipeline for prokaryotic and eukaryotic genomes
Abstract
Recommendation: posted 10 July 2022, validated 18 July 2022
Castanera, R. (2022) A flexible and reproducible pipeline for long-read assembly and evaluation. Peer Community in Genomics, 100018. https://doi.org/10.24072/pci.genomics.100018
Recommendation
Third-generation sequencing has revolutionised de novo genome assembly. Thanks to this technology, genome reference sequences have evolved from fragmented drafts to gapless, telomere-to-telomere genome assemblies. Long reads produced by Oxford Nanopore and PacBio technologies can span structural variants and resolve complex repetitive regions such as centromeres, unlocking previously inaccessible genomic information. Nowadays, many research groups can afford to sequence the genome of their working model using long reads. Nevertheless, genome assembly poses a significant computational challenge. Read length, quality, coverage and genomic features such as repeat content can affect assembly contiguity, accuracy, and completeness in almost unpredictable ways. Consequently, there is no best universal software or protocol for this task. Producing a high-quality assembly requires chaining several tools into pipelines and performing extensive comparisons between the assemblies obtained by different tool combinations to decide which one is the best. This task can be extremely challenging, as the number of tools available rises very rapidly, and thorough benchmarks cannot be updated and published at such a fast pace.
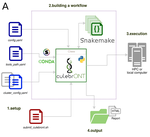
In their paper, Orjuela and collaborators present CulebrONT [1], a universal pipeline that greatly contributes to overcoming these challenges and facilitates long-read genome assembly for all taxonomic groups. CulebrONT incorporates six commonly used assemblers and allows to perform assembly, circularization (if needed), polishing, and evaluation in a simple framework. One important aspect of CulebrONT is its modularity, which allows the activation or deactivation of specific tools, giving great flexibility to the user. Nevertheless, possibly the best feature of CulebrONT is the opportunity to benchmark the selected tool combinations based on the excellent report generated by the pipeline. This HTML report aggregates the output of several tools for quality evaluation of the assemblies (e.g. BUSCO [2] or QUAST [3]) generated by the different assemblers, in addition to the running time and configuration parameters. Such information is of great help to identify the best-suited pipeline, as exemplified by the authors using four datasets of different taxonomic origins. Finally, CulebrONT can handle multiple samples in parallel, which makes it a good solution for laboratories looking for multiple assemblies on a large scale.
References
1. Orjuela J, Comte A, Ravel S, Charriat F, Vi T, Sabot F, Cunnac S (2022) CulebrONT: a streamlined long reads multi-assembler pipeline for prokaryotic and eukaryotic genomes. bioRxiv, 2021.07.19.452922, ver. 5 peer-reviewed and recommended by Peer Community in Genomics. https://doi.org/10.1101/2021.07.19.452922
2. Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics, 31, 3210–3212. https://doi.org/10.1093/bioinformatics/btv351
3. Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics, 29, 1072–1075. https://doi.org/10.1093/bioinformatics/btt086
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
Reviewed by Valentine Murigneux, 03 Jul 2022
I would like to thank the authors for answering all my comments and questions. I highly recommend the revised manuscript for publication.
https://doi.org/10.24072/pci.genomics.100018.rev21Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.1101/2021.07.19.452922
Version of the preprint: 3
Author's Reply, 24 Jun 2022
Decision by Raúl Castanera, posted 25 Apr 2022
Dear Authors,
Thank you for submitting your work to PCI genomics.
The two reviewers and I see your pipeline as a very useful tool for the community that will facilitate the production and evaluation of genome assemblies. The reviewers acknowledge the important number of tools included, the clear output summary report and the excellent documentation. They provide several suggestions on the text that I think will facilitate the understanding of the pipeline by the reader, as well as some minor technical comments to ease the installation process.
I suggest addressing the reviewer comments before I can recommend a revised version of your manuscript.
Sincerely,
Raúl Castanera
Reviewed by Benjamin Istace, 15 Mar 2022
I read the manuscript with great interest. The authors describe a new pipeline named "CulebrONT" that they developed in order to be able to test multiple genome assemblers at once. The pipeline also performs optional steps like the polishing and the circularization and outputs QC metrics that are often used to assess the quality of genome assemblies. I personally think that this type of pipeline is very useful to the community, as it aggregates the most commonly used tools in order to improve the ease of use for the end-user. I only have very minor concerns that I would like the authors to address if they agree with me.
Introduction - line 30-31: "Oxford Nanopore Technologies (ONT) and Pacific Biosciences (PB), provide reads up to hundreds of thousands of bases in length" - While I agree with this statement for ONT reads, PacBio reads are generally around 15kb and I don't think I have ever seen a read larger than 30kb. Would you have a reference for this?
I also rapidly tested CulebrONT on a small yeast genome and I also have some suggestions:
- R is not specified as a dependency but it is required to install the CulebrONT PyPI package (it is required by RPy2, which is a dependency of Pandas). I think that stating that it is required would be a good idea because it produces weird error messages otherwise.
- during the installation step (install_cluster), I chose to use the singularity environment. I think that it would be a good thing to indicate in the docs that images will be downloaded in CulebrONTs install directory. Indeed, it was installed in my home for testing purposes and completely filled it up. It was an easy fix to create a virtualenv in a more spacious filesystem but seeing it mentioned somewhere would be better I think.
Globally, the pipeline is relatively easy to use and configure with helpful messages.
Some general comments/suggestions, with no impact on the result of the review:
- the inclusion of Smartdenovo in the list of assemblers is a good point, as we often get good results with it but is a lesser-known software. You could also take a look at Necat, which is a new assembler that often leads to pretty great assemblies of complex genomes.
- I don't want to seem like I am pushing my own tool but for the polishing step with short reads, we developed Hapo-G which specifically handles heterozygous genomes while still doing great with homozygous/haploid ones. It's just a comment, I won't take it personally if it is not considered for this pipeline.
- Merqury is another tool that is very practical to assess the quality of an assembly. It is used with Illumina short reads and compares the k-mers that are in the assembly to the ones of the Illumina reads. It then gives a Q-score to the assembly based on shared k-mers.
- I am a big fan of Singularity and containers in general so seeing them included in a pipeline makes me very happy.
Reviewed by Valentine Murigneux, 20 Apr 2022
The manuscript describes the software tool culebrONT, whose goal is to help benchmark assembly pipeline. The introduction clearly explains the motivation of the pipeline development. This is a very useful tool that should be useful to many in the genome assembly community, who can be easily overwhelmed by a growing number of tools available and the fact that no tool performs best for every sample dataset. To my knowledge, there is no similar worklow/ software currently available in the community. The pipeline aims to solve common challenges for the user to install different tools prior to running them and comparing their results. Raw data and the source code are available to the reader. The pipeline is extremely well documented, illustrated and currently well maintained with an active Github webpage. A useful feature of the software is the Html report generated containing results, multiple graphes and the version of the tools.
I have a few questions and suggestions:
-line 14" Implementation
CulebrONT uses Snakemake [4] functionalities, enabling readability of the code,
local and HPC scalability, reentry, reproducibility and modularity. "
I am not familiar with snakemake functionalities therefore it could be useful to provide a few details on each aspect for the reader.
-Following up on the previous suggestion , I was looking for more details about the "modular" aspect of the pipeline. How easy is it for a user to add a new tool to the pipeline, e.g. a new assembler or polisher? Can a user do it thanks to the modular aspect of the pipeline and its open source status?
- Same question for a new version of a tool.
Can the user choose to use a new version of any tool, i.e. a more recent than the one listed on this page? https://culebront-pipeline.readthedocs.io/en/2.0.1/ABOUT.html#assembly
-the scalable aspect of the pipeline could be illustrated by a few examples. I wonder if examples could come from the "Application" section which contains several use cases from "personal communication" especially plants which requires more computational resources. Is it possible /useful to provide more details here.
-The manuscript does not contain a discussion section. The authors could comment on future developments/improvements planned for the pipeline if there are any. How and how often are the authors planning to maintain/ update/ improve the pipeline?
- The report includes the run time for each step of the pipeline. Is there an easy way for snakemake to also include the computational resources used e.g. memory/CPU ?
- Table 1: the legend does not mention if those examples are exclusively from ONT data?
-Table 1: the busco score for the nematode sample is quite low 65%, is there an explanation?
-The background section mentions past research in the field and available software. culebrONT aims at providing a workflow chaining different tools to facilitate genome assembly and compare different assembly results. Although restricted to prokaryotic genomes, previous benchmarkings of long read assemblers could be cited in the introduction (e.g. https://f1000research.com/articles/8-2138, https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7730629/, https://www.nature.com/articles/s41598-020-70491-3) as well as a workflow for bacterial genome assembly using long read sequencing published in 2021 (https://bmcgenomics.biomedcentral.com/articles/10.1186/s12864-021-07767-z). CulebrONT includes a lot of similar tools as included in those publications. CulebrONT provides the advantages of reporting the results of several combination of tools to facilitate their comparison.
https://doi.org/10.24072/pci.genomics.100158.rev12









